Visualization
This is a list of analysis and quantification software.
CloudBrain-MRS
CloudBrain-MRS is a cloud-based computational platform for MRS data. The platform deploys the mainstream quantification tool LCModel and advanced deep learning denoising and quantification algorithms. The platform has been developed with a module for the statistical analysis of biomarkers. Users can batch preprocess and quantitative analysis MRS data with only one browser. The platform has been developed with a range of visualization techniques to provide a user-friendly interface.
| CloudBrain-MRS | |
|---|---|
| Developer | Jiayu Li, Xiaodie Chen, Dicheng Chen, Yirong Zhou, Zhangren Tu, Meijin Lin, Xiaobo Qu |
| Language | Python, JavaScript |
| License | BSD3 |
| Credit | Please reference the following publication if you use CloudBrain-MRS: X. Chen et al., “CloudBrain-MRS: An intelligent cloud computing platform for in vivo magnetic resonance spectroscopy preprocessing, quantification, and analysis,” arXiv:2306.11021, 2023. D. Chen et al., “Magnetic resonance spectroscopy deep learning denoising using few in vivo data,” IEEE Trans. Comput. Imaging, vol. 9, pp. 448-458, 2023. D. Chen et al., “Magnetic resonance spectroscopy quantification aided by deep estimations of imperfection factors and overall macromolecular signal,” arXiv:2306.09681, 2023. |
VDI

VDI (Visual Display Interface) is a collection of MATLAB libraries created to visualize and process multiresolution data, such as MRS, MRSI, fMRI and anatomical MRI. An additional sub-library called SpinTool (independently available for download) can be used to easily and quickly write Bloch simulations and simulations of J-coupled spins.
| VDI | |
|---|---|
| Developer | Assaf Tal |
| Language | MATLAB |
| License | BSD3 |
| Credit | Please cite the software package using the APS style or equivalent (Tal, A (2020). Visual Display Interface (VDI) [Computer Software]. Retrieved from http://www.vdisoftware.net) |
FSL-MRS
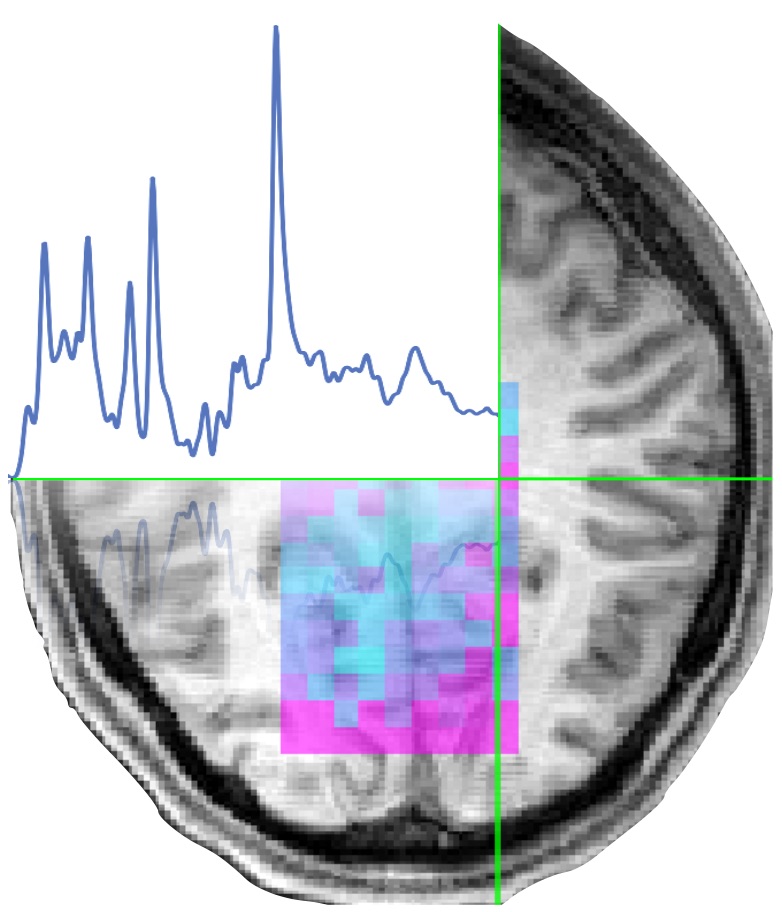
FSL-MRS is a collection of python modules and wrapper scripts for pre-processing and model fitting of Magnetic Resonance Spectroscopy (MRS) and Spectroscopic Imaging (MRSI) data.
| FSL-MRS | |
|---|---|
| Developer | William Clarke and Saad Jbabdi |
| Language | Python |
| License | FSL License https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/Licence |
| Credit | If you use FSL-MRS in your research please cite the publication below. |
MRS-voxel-plot
Simple scripts to produce informative MRS voxel and spectral plots for a group or groups of participants.
| MRS-voxel-plot | |
|---|---|
| Developer | Niall Duncan, Vuong Truong |
| Language | Python |
| License | MIT |
| Credit | Please cite the publication mentioned below if you use this code. |
MRSHub Code Author Website Publication
Oryx-MRSI

ORYX-MRSI: A data analysis software for multi-slice 1H-MRSI
| Oryx-MRSI | |
|---|---|
| Developer | Sevim Cengiz |
| Language | MATLAB |
| License | MIT |
| Credit | Please cite the following publication if you use Oryx-MRSI: Cengiz S, Yildirim M, Bas A, Ozturk-Isik E. ORYX-MRSI: A data analysis software for multi-slice 1H-MRSI. International Society for Magnetic Resonance in Medicine. Virtual Meeting, May 15-20, 2021 (Digital Poster) |
Osprey

Osprey is an all-in-one open-source software suite for state-of-the art processing, quantitative analysis, and visualization of in-vivo magnetic resonance spectroscopy (MRS) data.
| Osprey | |
|---|---|
| Developer | Georg Oeltzschner, Helge J Zöllner, Richard AE Edden |
| Language | MATLAB |
| License | MIT |
| Credit | Should you publish material that made use of Osprey, please cite the publication below. |
pysoher
A collection of algorithms and general code tools (GUI, I/O etc.) for use with MRS research.
| pysoher | |
|---|---|
| Developer | Brian J. Soher |
| Language | Python |
| License | BSD |
| Credit | BSD license, byproducts of the Vespa Versatile Simulation Pulses and Analysis project for MRS research at https://scion.duhs.duke.edu/vespa/project |
QMRITools

QMRITools is written in Mathematica using Wolfram Workbench and Eclipse and contains a collection of tools and functions for processing quantitative MRI data. The software tools for spectroscopy are mostly focussed on 31P CSI analysis but do not exclude other nuclei and/or acquistion methods. The basic features include methods for manipulating and processing spectra (including PCA denoising and deconvolution), generation of basis spectra using j-coupling simulations and fitting of basis spectra to the spectra.
| QMRITools | |
|---|---|
| Developer | Martijn Froeling, PhD |
| Language | Mathematica |
| License | BSD-3 |
| Credit | Please cite the publication mentioned below if you use QMRITools. A publication on the spectro specific tools is in preparation. |
Author Website Publication Publication 2 Publication 3
SIVIC

SIVIC is an open-source, standards-based software framework and application suite for processing and visualizing DICOM MR Spectroscopy data.
| SIVIC | |
|---|---|
| Developer | Nelson Lab, UCSF |
| Language | C++ |
| License | Open-source, https://github.com/SIVICLab/sivic/blob/master/LICENSE |
| Credit | |
SMAL
The Stanford CNI MRS Library (SMAL) provides algorithms and methods to read and analyze data from Magnetic Resonance Spectroscopy (MRS) experiments. It provides an API for fitting models of the spectral line-widths of several different molecular species, and quantify their relative abundance in human brain tissue
| SMAL | |
|---|---|
| Developer | Ariel Rokem, PhD |
| Language | Python |
| License | BSD3 |
| Credit | |
SpecVis

SpecVis is a repository of R functions to visualize quantitative MRS results from different linear-combination algorithms.
| SpecVis | |
|---|---|
| Developer | Helge J. Zöllner, PhD |
| Language | R, MATLAB |
| License | MIT |
| Credit | Please cite the publication mentioned below if you use SpecVis (Zöllner et al, Comparison of algorithms for linear-combination modelling of short-echo-time magnetic resonance spectra, bioRxiv (2020)). |