MRSI datasets
This is a list of MRSI datasets.
Inter-subject stability and regional concentration estimates of 3D-FID-MRSI in the human brain at 7T
Intended as supplementary data to the manuscript ‘Inter-subject stability and regional concentration estimates of 3D-FID-MRSI in the human brain at 7T’ It contains five full volunteer datasets in MINC and NIFTI format.
| Inter-subject stability and regional concentration estimates of 3D-FID-MRSI in the human brain at 7T | |
|---|---|
| Developer | Gilbert Hangel, Benjamin Spurny, Philipp Lazen, Cornelius Cadrien, Sukrit Sharma, Lukas Hingerl, Eva Hečková, Bernhard Strasser, Stanislav Motyka, Alexandra Lipka, Stephan Gruber, Christoph Brandner, Rupert Lanzenberger, Karl Rössler, Siegfried Trattnig, Wolfgang Bogner |
| Format | NIfTI, MINC |
| Sequence | FID-MRSI |
| License | creative commons attribution 4.0 international license |
| Credit | Hangel et al. 2020-21 |
| Contact | gilbert.hangel@meduniwien.ac.at |
FID-MRSI 3T and 7T basis sets with measured MM
These are FID-MRSI basis sets with metabolite information simulated in NMR-Scope B and measured MM (see Publication Povazan et al, NI, 2015)
| FID-MRSI 3T and 7T basis sets with measured MM | |
|---|---|
| Developer | Wolfgang Bogner, Gilbert Hangel, Michal Povazan, Bernhard Strasser |
| Format | LCModel .basis and .RAW |
| Sequence | FID-MRSI with acquisition delay of 0 ms / 1.3 ms / 1.5 ms |
| License | BSD3 |
| Credit | Please consult the authors how to cite these basis sets |
| Contact | mpovaza1@jhmi.edu; wolfgang.bogner@meduniwien.ac.at |
7T-CRT-FID-MRSI (HFMRC, MU Vienna) Example Data - Raw Spectra
Data from the Vienna 7T MRSI group. Combined and preprocessed data after spatial FFT, in the time domain. 64x64x39 matrix, 15 min. Method details: Hingerl et al 2020, doi: 10.1097/RLI.0000000000000626. Healthy_Volunteer.mat: Female, 24y, measured 2019. GBM_patient.mat: Glioblastoma WHO Grade 4, with IDH1 mutation, male, 52y.
| 7T-CRT-FID-MRSI (HFMRC, MU Vienna) Example Data - Raw Spectra | |
|---|---|
| Developer | Wolfgang Bogner, Gilbert Hangel |
| Format | .MAT |
| Sequence | CRT-FID-MRSI (7T) |
| License | BSD3 |
| Credit | |
| Contact | wolfgang.bogner@meduniwien.ac.at; gilbert.hangel@meduniwien.ac.at |
MRSI Data for 'High-Resolution Metabolic Imaging of High-Grade Gliomas using 7T-CRT-FID-MRSI'
MRSI Maps from the Vienna 7T scanner in MINC format. Intended as supplementary data to the manuscript ‘High-Resolution Metabolic Imaging of High-Grade Gliomas using 7T-CRT-FID-MRSI’ (Authors: Gilbert Hangel, Cornelius Cadrien, Philipp Lazen, Julia Furtner, Alexandra Lipka, Eva Hečková, Lukas Hingerl, Stanislav Motyka, Stephan Gruber, Bernhard Strasser, Barbara Kiesel, Mario Mischkulnig, Matthias Preusser, Thomas Roetzer, Adelheid Wöhrer, Georg Widhalm, Karl Rössler, Siegfried Trattnig and Wolfgang Bogner). For use with the minc-toolkit: https://bic-mni.github.io/. MRSI method published as Hingerl et al 2020, doi: 10.1097/RLI.0000000000000626. Patient 1: Glioblastoma WHO Grade 4, with IDH1 mutation, male. Patient 2: Glioblastoma WHO Grade 4, with IDH1 mutation, male.
| MRSI Data for 'High-Resolution Metabolic Imaging of High-Grade Gliomas using 7T-CRT-FID-MRSI' | |
|---|---|
| Developer | Wolfgang Bogner, Gilbert Hangel |
| Format | MINC |
| Sequence | CRT-FID-MRSI (7T) |
| License | BSD3 |
| Credit | |
| Contact | wolfgang.bogner@meduniwien.ac.at; gilbert.hangel@meduniwien.ac.at |
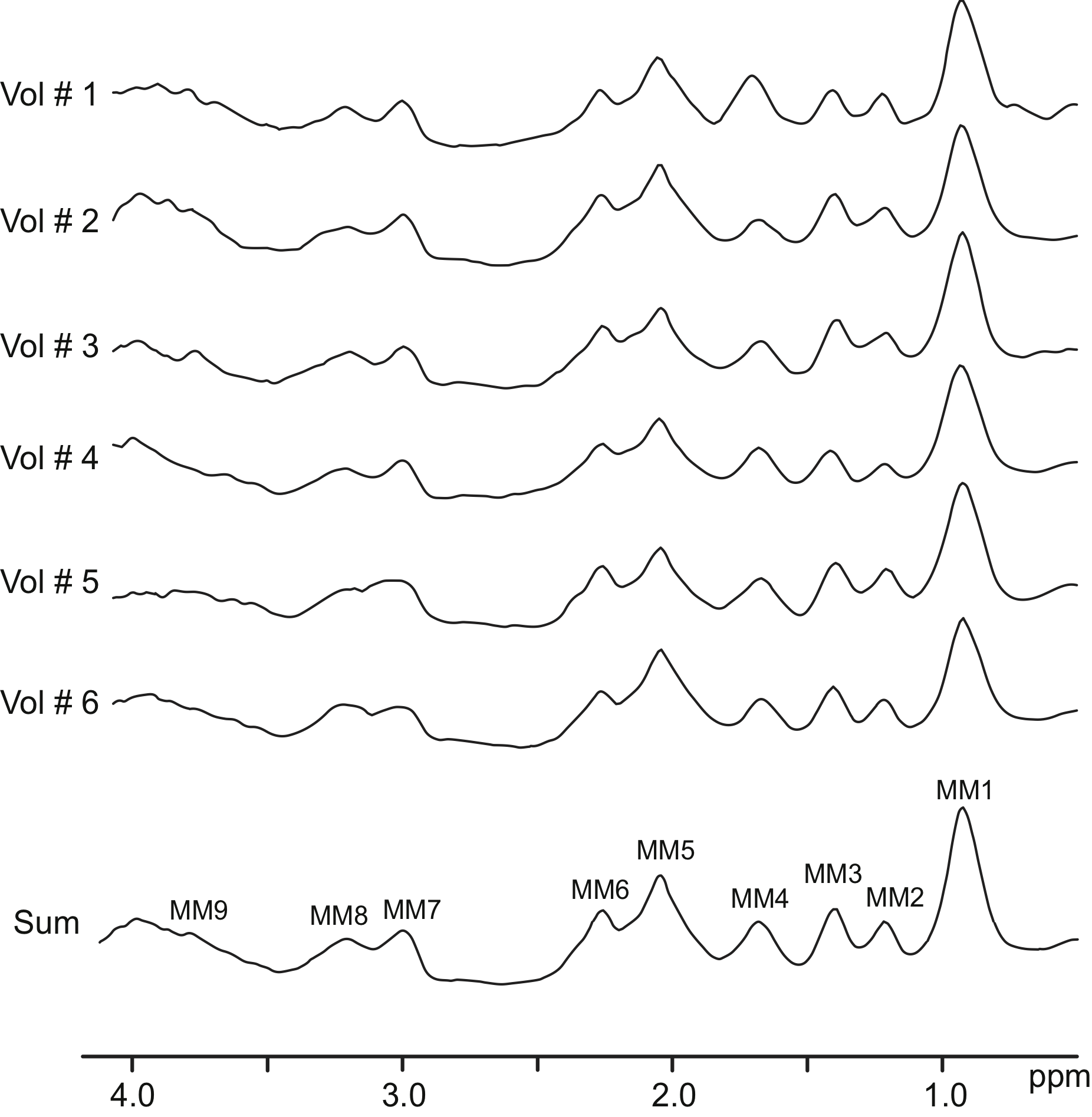
7T-FID-MRSI example of metabolite-nulled macromolecules spectra

This is the example of metabolite-nulled macromolecules spectra acquired from 6 healthy young volunteers scanned at 7T using double-inversion recovery FID-MRSI
| 7T-FID-MRSI example of metabolite-nulled macromolecules spectra | |
|---|---|
| Developer | Michal Povazan, Wolfgang Bogner |
| Format | jMRUI .txt |
| Sequence | double-inversion recovery single slice FID-MRSI |
| License | BSD3 |
| Credit | Please cite Považan M, Hangel G, Strasser B, Gruber S, Chmelik M, Trattnig S, et al. Mapping of brain macromolecules and their use for spectral processing of 1H-MRSI data with an ultra-short acquisition delay at 7T. Neuroimage. 2015;121:126–35if you use this dataset. |
| Contact | mpovaza1@jhmi.edu; wolfgang.bogner@meduniwien.ac.at |